I am working my way through the Udemy data science course “Machine Learning A-Z” by Kirill Eremenko and Hadelin de Ponteves. The course steps through key machine learning algorithms and approaches using Python and R. As an R programmer, it’s great to compare to the Python code and learn it’s syntax. From my nascent observations, it takes fewer lines to code an approach with R compared to Python.
Kirill and Hadelin are clear communicators. They break-down complex information, guiding the viewer with palatable bite-sized chucks of information. I was so impressed that I sent Kirill a thank you on Udemy. Kirill responded, we added each other on LinkedIn, then he invited me as a guest on his podcast at Super DataScience!
My episode can be found here, here and here. Three links, same episode - Woo!
Thanks to Kirill for having me as a guest and giving me an excuse to talk about neuroscience – something I haven’t done for the past three years. The dorsal lateral prefrontal cortex got a mention :)
Showing posts with label R. Show all posts
Showing posts with label R. Show all posts
Wednesday, 9 November 2016
Tuesday, 13 September 2016
Building plots with ggraptR’s code gen
Building
plots for R newbies is a challenge, even for R not-so-newbies like myself. Why
write code when it can be generated for you?
I have put my hand up to volunteer towards an R visualisation package called ggraptR. ggraptR allows interactive data visualisation via a web browser GUI (demonstrated in a previous post using my Fitbit data). The latest Github version (as of 13th September 2016) contains a plotting code generation feature. Let’s take it for a spin!
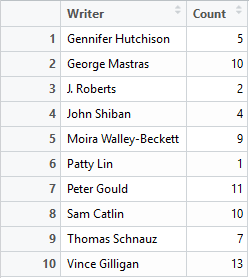
I have a rather simple data frame called ""dfGroup" that contains the number of Breaking Bad episodes each writer wrote. I want to create a horizontal bar plot with the “Count” on the x-axis and “Writer” on the y-axis. The writers will be ordered from most episodes written (with Mr Vince Gillian at the top) to least (bottom). It will have an awesome title and awesomely-labelled axis. The bars will be green. Breaking Bad green.
With a tiny bit of modifying – adding a title, changing the axis titles and filling in the bars with Breaking Bad green, we have the following:
Using ggraptR you can quickly build a plot, use code gen to copy the code then modify it as desired. Happy plotting!
I have put my hand up to volunteer towards an R visualisation package called ggraptR. ggraptR allows interactive data visualisation via a web browser GUI (demonstrated in a previous post using my Fitbit data). The latest Github version (as of 13th September 2016) contains a plotting code generation feature. Let’s take it for a spin!
I have a rather simple data frame called ""dfGroup" that contains the number of Breaking Bad episodes each writer wrote. I want to create a horizontal bar plot with the “Count” on the x-axis and “Writer” on the y-axis. The writers will be ordered from most episodes written (with Mr Vince Gillian at the top) to least (bottom). It will have an awesome title and awesomely-labelled axis. The bars will be green. Breaking Bad green.
Before code gen, I would Google “R horizontal bar ggplot
with ordered bars”, copy paste code then adjust it by adding more code. The
ggraptR approach begins with installing and loading the latest build:
devtools::install_github('cargomoose/raptR', force = TRUE)
devtools::install_github('cargomoose/raptR', force = TRUE)
library("ggraptR")
Launch ggraptR with ggraptR().
Launch ggraptR with ggraptR().
A
web browser will launch. Under “Choose a dataset” I selected my dfGroup data
frame. Plot Type is “Bar”. The selected X axis is “Writer” and the Y is
“Count”. “Flip X and Y coordinates” is checked. And voilà – instant horizontal
bar plot.
Notice
the “Generate Plot Code” button highlighted in red. Clicking on said button – a
floating window with code will appear.
I copied and pasted the code in an R script. I tidied the code a bit as shown below. Running the code (with dfGroup in the environment) will produce the plot as displayed with ggraptR.
One last thing – the bars are not ordered. Currently the
bars cannot be ordered with ggraptR. I can reorder the bars using the reorder
function on the dfGroup data frame. Back in RStudio, I run the following:
dfGroup$Writer <- reorder(dfGroup$Writer, dfGroup$Count)
then execute the modified code above and we have plotting
success!
Sunday, 29 May 2016
Fitbit 03 – Getting and wrangling all data
Previous post in this series: Fitbit 02 – Getting and wrangling sleep data.
This post will wrap-up the getting and wrangling of Fitbit
data using fitbitscraper.
This is the list of data that was gathered [1]:
- Steps
- Distance
- Floors
- Very active minutes (“MinutesVery”)
- Calories burned
- Resting heart rate (“RestingHeart”)
- Sleep
- Weight.
For each dataset, the data was gathered then wrangled as
separate tidy data frames. Each data contained a unique date per row. Most
datasets required minimal wrangling. A previous post outlined the extra effort
required to wrangle sleep data due to split sleep sessions and some extra looping to gather all weight data.
Each data frame contains a Date column. The data frames are
joined by the unique dates to create one big happy data frame of Fitbitness.
Each row is a date containing columns of fitness factors.
Now what? I feel like a falafel. I’m going to eat a falafel
[2].
With this tidy dataset I will continue the analytics journey
in future posts. For now, I wish to quickly visualise the data. Writing lines
of code for plots in R is not-so-quick. Thankfully there’s a point-and-click
visualisation package available called ggraptR. Installing and launching the
package is achieved as follows.
devtools::install_github('cargomoose/raptR', force = TRUE) # install
library("ggraptR") # load
ggraptR() # launch
My main hypothesis was that steps/distance may correlate
with weight. There was no relationship observed on a scatter plot. This is
preliminary, future post will focus on exploratory data analysis. Prior to data
analysis I need to ask some driving questions.
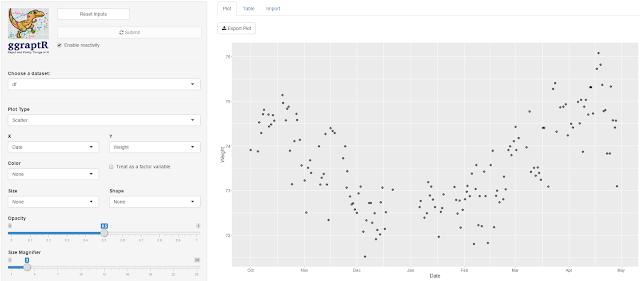
I plotted Date vs Weight. My weight fell gradually from
October 2015 through to December. I was on a week-long Sydney to Adelaide road
trip during the end of December, got a parking ticket in Adelaide and did not
have recorded weights whilst on the road. My weight steadily increased since.
Not a lot of exercise, quite a lot of banana Tim Tams.
After sequential pointing-and-clicking, I overlayed this
time plot with another factor - the “AwakeBetweenDuration”. In the previous
post I noted I wake-up in the middle of the night. It may take hours before I
fall asleep again. The tidy dataset holds the number of minutes awake between
such sessions. The bigger the bubble, the longer I was awake between sleep sessions.
Here’s a driving question: what accounts for the nights when
I am awake for long durations? I was awake some nights in October, December
(some of my road trip nights – I couldn’t drive for one of those days as I was
exhausted), January and then April. February and March appeared almost
blissful. Why? Tell me data, why?
Here is the Fitbit data wrangling code published on GitHub,
FitbitWrangling.R: https://github.com/muhsinkarim/fitbit Replace “your_email”
and “your_password” with your email and your password used to log into your
Fitbit account and dashboard.
References and notes
1. The fitbitscraper
function get_activity_data() will return rows of activities per day including
walking and running. I only have activity data from 15th February
2016. Since I’m analysing data since October 2015 (where I have weight data
from my Fitbit scales) I chose not in include activity data in the tidy
dataset.
2. I ate two.
Sunday, 15 May 2016
Fitbit 02 – Getting and wrangling sleep data
Previous post in this series: Fitbit 01 – Getting and wrangling weight, steps and calories burned data.
Let’s continue to get and wrangle Fitbit data. This post will tackle sleep data and address my unattractive trait of not easily letting things go.
I applied the following R package fitbitScraper function wrapped in a data frame:
There’s a lot of data. Below is the list of columns I want to keep:
If you’re a normalish human being, you would look at the sleep data and examine it for any anomalies. Your keen eye will note that some sleep durations are split across two or more sessions. That is, you went to bed, woke up in the middle of the night, then fell asleep again. Depending on how long you were awake for, Fitbit will record separate sessions.
I am working towards a tidy dataset with each row representing a unique date with the day’s Fitbit data. Separate sleep sessions cause duplicate dates. Being a normalish human being, you write code that will group the split sleep sessions back together. Unique dates per sleep lends to tidy data.
If you’re not a normal human being, you examine the intricate nature of these split sleep sessions and spend way too much time writing code that groups the data back together.
I am not a normal human being.
Split sessions occur for two reasons. The first occurs when I wake up in the early hours of the morning. An example is below.
There’s a final consideration with split sleep sessions. Consider the below.
Let’s continue to get and wrangle Fitbit data. This post will tackle sleep data and address my unattractive trait of not easily letting things go.
I applied the following R package fitbitScraper function wrapped in a data frame:
dfSleep <- as.data.frame(get_sleep_data(cookie, start_date = startDate, end_date = endDate))
There’s a lot of data. Below is the list of columns I want to keep:
- df.startDateTime – The start datetime of sleep.
- df.endDateTime – The end datetime of sleep.
- df.sleepDuration – The minutes between the start and end time. df.sleepDuration = df.awakeDuration + df.restlessDuration + df.minAsleep".
- df.awakeDuration – The minutes of wakefulness.
- df.restlessDuration – The minutes of restlessness.
- df.minAsleep – The minutes of actual sleep. I want to maximise this data field. More sleep less grumpiness.
If you’re a normalish human being, you would look at the sleep data and examine it for any anomalies. Your keen eye will note that some sleep durations are split across two or more sessions. That is, you went to bed, woke up in the middle of the night, then fell asleep again. Depending on how long you were awake for, Fitbit will record separate sessions.
I am working towards a tidy dataset with each row representing a unique date with the day’s Fitbit data. Separate sleep sessions cause duplicate dates. Being a normalish human being, you write code that will group the split sleep sessions back together. Unique dates per sleep lends to tidy data.
If you’re not a normal human being, you examine the intricate nature of these split sleep sessions and spend way too much time writing code that groups the data back together.
I am not a normal human being.
Split sessions occur for two reasons. The first occurs when I wake up in the early hours of the morning. An example is below.
On the night of the 22nd March I fell asleep at
23:37. I woke up at 3:49 on the 23rd. After hating my life and eating
morning chocolate, I finally fell asleep again at 4:51. Technically the second
session date should be displayed as the 23rd of March, not the 22nd.
These display dates are akin to the “date I tried to fall asleep”. The
datetimes record the true date and times.
When I combine these separate sessions, the new sleep start
time will be 23:37 and the new sleep end time will be 07:46. The SleepDuration,
SleepAwakeDuration, SleepRestlessDuration, SleepMinAsleep, SleepAwakeCount and
SleepRestlessCount values will be summed together. I would also like to note
the number of minutes I spent awake between sessions and the number of separate
sessions (two in this example).
The second split sleep session type appears to be a glitch with
sessions being separated by a difference of one minute. An example is below.
On the 23rd of February I have sleep sessions ending
at 03:47 then resuming at 03:38. There are multiple instances where this occurs
in my datasets. As before, the sleep variables from both sessions will be
summed. I don’t need to note the minutes between the sessions as the one minute
difference is meaningless. Further, the number of sleep sessions should be
recorded as one, not two.
According this this display, on the 8th April I slept from 22:23 to 5:27, then on the 21:06 to the 23:57 on the same day. No I didn’t! The sleep session datetimes are overlapping. The session displayed on the 8th April from 21:06 to 23:57 should read the 9th April, not the 8th. This session needs to be combined with the sessions displayed on the 9th April from the 23:58 to 6:03. I may not be a normal human being, but I have my limits. For this last case, I did not write a patch of code that could group the data appropriately. Since there were few instances when such overlapping sessions occurred, I let it go – such sessions were removed from the dataset resulting in missing sleep data for that particular date.
Here is a quick plot of the average number of minutes asleep
per weekday. Nothing out of the ordinary. I sleep an average 7.6 hours on
Sunday nights and an average 6.4 hours on Wednesday nights. I can’t think of a
reason why I get fewer hours on a Wednesday night.
I spend most of my time wrangling data. I come across
problems in datasets as described above often, and write code to return the
numbers back to reality as much as possible. When the costs (time and effort) outweigh
the benefits (more clean data) I have to let some data go and remove it.
I have outlined the code at the end of this post for the
avid reader.
#### Sleep
### Get data
dfSleep <- as.data.frame(get_sleep_data(cookie, start_date = startDate, end_date = endDate))
### Keep key columns
dfSleep <- dfSleep[ , c("df.date", "df.startDateTime", "df.endDateTime", "df.sleepDuration", "df.awakeDuration",
"df.restlessDuration", "df.minAsleep")]
## Rename colnames
# Date is sleep date attempt
colnames(dfSleep) <- c("Date","SleepStartDatetime", "SleepEndDatetime", "SleepDuration",
"SleepAwakeDuration", "SleepRestlessDuration", "SleepMinAsleep")
### Combine the split sleep sessions
## Index the Dates that are duplicated along with their original
duplicatedDates <- unique(dfSleep$Date[which(duplicated(dfSleep$Date))])
dfSleep$Combine <- ""
dfSleep$Combine[which(dfSleep$Date %in% duplicatedDates)] <-
dfSleep$Date[which(dfSleep$Date %in% duplicatedDates)]
## Subset the combine indexed rows
dfSubset <- dfSleep[which(dfSleep$Combine != ""), ]
## Aggregate rows marked to combine
dfSubset <-
dfSubset %>%
group_by(Combine) %>%
summarise(Date = unique(Date),
SleepStartDatetime = min(SleepStartDatetime), # Earliest datetime
SleepEndDatetime = max(SleepEndDatetime), # Latest datetime
SleepDuration = sum(SleepDuration),
SleepAwakeDuration = sum(SleepAwakeDuration),
SleepRestlessDuration = sum(SleepRestlessDuration),
SleepMinAsleep = sum(SleepMinAsleep),
SleepSessions = n() # Number of split sleep sessions
)
### Get the minutes awake between split sessions
## Calculate sleep duration using start and end time using floor (round down)
dfSubset$AwakeBetweenDuration <- floor((as.POSIXct(dfSubset$SleepEndDatetime) - as.POSIXct(dfSubset$SleepStartDatetime)) * 60)
dfSubset$AwakeBetweenDuration <- dfSubset$AwakeBetweenDuration - dfSubset$SleepDuration
## Set any duration less than five minutes to zero
dfSubset$AwakeBetweenDuration[which(dfSubset$AwakeBetweenDuration < 5)] <- 0
### Remove rows of combined sleep sessions
## Remove any row with AwakenBetweenDuration greater than five hours
dfSubset <- dfSubset[-which(dfSubset$AwakeBetweenDuration > 60*5), ]
### Replace split sessions in dfSleep with dfSubset
## Prepare dfSubset for merging
## Remove duplicate dates from dfSleep
dfSleep <- dfSleep[-which(nchar(dfSleep$Combine) > 0), ]
## Add new columns
dfSleep$SleepSessions <- 1
dfSleep$AwakeBetweenDuration <- 0
## Remove Combine column
dfSubset <- dfSubset[ , -which(colnames(dfSubset) == "Combine")]
dfSleep <- dfSleep[ , -which(colnames(dfSleep) == "Combine")]
## Bind dfSubset
dfSleep <- rbind.data.frame(dfSleep, dfSubset)
### Coerce as date
dfSleep$Date <- as.Date(dfSleep$Date)
Sunday, 24 April 2016
Fitbit 01 – Getting and wrangling weight, steps and calories burned data
I love my Fitbit Charge HR [1]. It tracks my steps, it tracks my heartbeat. It tracks my sleep hours. Alas it does not keep me warm at night unless if malfunctions, bursts into flames then rudely singes my beard.
I’m keen to analyse my Fitbit data to gain some insight into my biology and behaviour. What days and times am I the most active? What drives my weight gain and loss? What do I need to do to get a decent night’s sleep? I will attempt to answer such questions through a series of Fitbit posts, each one taking a snapshot of the process from data to insights. This post is focussed on getting the Fitbit data, then wrangling it into tidy data for further visualisation and analysis.
Accessing my Fitbit data is made easy with the R package fitbitScraper. I have been using my Fitbit since March 2015. I have been recording my weight via the Fitbit Aria Wi-Fi Smart Scale since the end of September 2015. I will analyse data from October 2015 (a complete month since recording my weight) to March 2016 (seven months). These are the data variables of interest:
I will focus on weight, steps and calories burned for this post.
Weight
After authenticating [2], assigning my start and end date, I applied the get_weight_data function and received the following output:
I have more weights recorded which are not being captured. I tried setting the dates within a single month (March 2016) and it returned the following:
Huzzah! All my March weights are visible (as verified by checking against the Fitbit app). I wrote code that loops through each month and binds all weight data. I will make this available on Github soonish. I noted duplicate dates in the data frame. That is, on some days I recorded my weight twice of a given day. As this analysis will focus on daily Fitbit data, I must have unique dates per variable prior to merging all the variables together. I removed the duplicate dates, keeping the weight recorded later on a given day since I tend to weigh myself at night. The final data frame contains two columns: Date (as class Date, not POSIXct) and Weight. Done.
Steps
Getting daily steps is easy with the get_daily_data function.
The date was coerced to class Date, columns renamed and that’s it.
Calories burned
Using the same get_daily_data function, I got the calories burned and intake data.
As with the steps, the date was coerced to class Date and the colnames renamed. The function returns both the calories burned each day and the intake of calories. Calories intake is gathered items entered into the food log. I only recently stopped recording my food. For items that did not have a barcode or were easily identifiable in the database, I would resort to selecting the closest match and guestimating serving sizes. I was underestimating how much I consumed each day as I was often below my target calories intake, yet I gained weight. I stopped recording food and will wait for a sensor to be surgically embedded in my stomach that quantifies the calories I shove down there. I disregarding the calories intake variable.
Merging the data
The data frames for weight, steps and calories burned were merged using Date.

There’s an obvious relationship since I burn calories with each step. Most of my activities involve making steps – walking, jogging, getting brownies, fleeing swooping birds. I do not clock-up steps when I kayak. I don’t know how Fitbit treats such activities. I’ll check the calorie count before and after next time.
Ultimately I’d like to observe whether any variables can account for outcomes such as weight and sleep. Here’s a Steps vs Weight plot.

There’s no relationship. It remains to be observed whether the inclusion of other Fitbit predictors could account for weight. More on this in future posts.
References and notes
1. I am not affiliated with Fitbit. I’m just a fan. I will however accept gifted Fitbit items if you would like to get in touch with me at willselloutfortech@gmail.com
2. Authenticate with the login function; login(email, password, rememberMe = FALSE). Use your email address and password you use to access your Fitbit dashboard online.
I’m keen to analyse my Fitbit data to gain some insight into my biology and behaviour. What days and times am I the most active? What drives my weight gain and loss? What do I need to do to get a decent night’s sleep? I will attempt to answer such questions through a series of Fitbit posts, each one taking a snapshot of the process from data to insights. This post is focussed on getting the Fitbit data, then wrangling it into tidy data for further visualisation and analysis.
Accessing my Fitbit data is made easy with the R package fitbitScraper. I have been using my Fitbit since March 2015. I have been recording my weight via the Fitbit Aria Wi-Fi Smart Scale since the end of September 2015. I will analyse data from October 2015 (a complete month since recording my weight) to March 2016 (seven months). These are the data variables of interest:
- Weight
- Sleep
- Steps
- Distance
- Activity
- Calories burned
- Heartbeat.
I will focus on weight, steps and calories burned for this post.
Weight
After authenticating [2], assigning my start and end date, I applied the get_weight_data function and received the following output:
> get_weight_data(cookie, start_date = startDate, end_date = endDate)
time weight
1 2015-09-27 23:59:59 72.8
2 2015-10-04 23:59:59 73.6
3 2015-10-11 23:59:59 74.5
4 2015-10-18 23:59:59 74.5
5 2015-10-25 23:59:59 74.5
6 2015-11-01 23:59:59 74.0
7 2015-11-08 23:59:59 73.3
8 2015-11-15 23:59:59 73.4
9 2015-11-22 23:59:59 74.3
10 2015-11-29 23:59:59 73.1
11 2015-12-06 23:59:59 72.3
12 2015-12-13 23:59:59 72.3
13 2015-12-20 23:59:59 72.5
14 2015-12-27 23:59:59 73.0
15 2016-01-10 23:59:59 72.6
16 2016-01-17 23:59:59 72.8
17 2016-01-24 23:59:59 72.7
18 2016-01-31 23:59:59 72.5
19 2016-02-07 23:59:59 72.8
20 2016-02-14 23:59:59 72.7
21 2016-02-21 23:59:59 73.1
22 2016-02-28 23:59:59 73.5
23 2016-03-06 23:59:59 74.0
24 2016-03-13 23:59:59 73.8
25 2016-03-20 23:59:59 73.7
26 2016-03-27 23:59:59 75.0
27 2016-04-03 23:59:59 74.6
28 2016-04-10 23:59:59 74.6
> get_weight_data(cookie, start_date = "2016-03-01", end_date = "2016-03-31")
time weight
1 2016-02-29 20:46:18 74.1
2 2016-03-01 21:04:14 74.4
3 2016-03-02 07:24:03 73.7
4 2016-03-02 21:10:52 73.9
5 2016-03-03 21:55:57 74.2
6 2016-03-08 20:09:37 74.0
7 2016-03-09 22:19:34 74.8
8 2016-03-10 20:06:33 73.4
9 2016-03-12 20:40:28 73.5
10 2016-03-13 21:03:40 73.3
11 2016-03-14 21:02:22 73.5
12 2016-03-15 22:13:07 73.4
13 2016-03-16 18:54:18 73.2
14 2016-03-17 21:02:32 74.4
15 2016-03-18 20:27:08 74.4
16 2016-03-20 18:21:45 73.1
17 2016-03-23 21:54:01 75.3
18 2016-03-24 20:03:48 75.4
19 2016-03-25 14:09:23 74.3
20 2016-03-27 20:18:27 74.9
21 2016-03-29 21:02:56 74.9
22 2016-03-30 22:08:01 74.9
23 2016-03-31 22:22:25 74.2
24 2016-04-02 22:29:36 74.5
Huzzah! All my March weights are visible (as verified by checking against the Fitbit app). I wrote code that loops through each month and binds all weight data. I will make this available on Github soonish. I noted duplicate dates in the data frame. That is, on some days I recorded my weight twice of a given day. As this analysis will focus on daily Fitbit data, I must have unique dates per variable prior to merging all the variables together. I removed the duplicate dates, keeping the weight recorded later on a given day since I tend to weigh myself at night. The final data frame contains two columns: Date (as class Date, not POSIXct) and Weight. Done.
Steps
Getting daily steps is easy with the get_daily_data function.
> dfSteps <- get_daily_data(cookie, what = "steps", startDate, endDate)
> head(dfSteps)
time steps
1 2015-10-01 7496
2 2015-10-02 7450
3 2015-10-03 4005
4 2015-10-04 2085
5 2015-10-05 3101
6 2015-10-06 10413
The date was coerced to class Date, columns renamed and that’s it.
Calories burned
Using the same get_daily_data function, I got the calories burned and intake data.
> dfCaloriesBurned <- get_daily_data(cookie, what = "caloriesBurnedVsIntake", startDate, endDate)
> head(dfCaloriesBurned)
time caloriesBurned caloriesIntake
1 2015-10-01 2428 2185
2 2015-10-02 2488 1790
3 2015-10-03 2353 2361
4 2015-10-04 2041 1899
5 2015-10-05 2213 2217
6 2015-10-06 4642 2474
As with the steps, the date was coerced to class Date and the colnames renamed. The function returns both the calories burned each day and the intake of calories. Calories intake is gathered items entered into the food log. I only recently stopped recording my food. For items that did not have a barcode or were easily identifiable in the database, I would resort to selecting the closest match and guestimating serving sizes. I was underestimating how much I consumed each day as I was often below my target calories intake, yet I gained weight. I stopped recording food and will wait for a sensor to be surgically embedded in my stomach that quantifies the calories I shove down there. I disregarding the calories intake variable.
Merging the data
The data frames for weight, steps and calories burned were merged using Date.
> df <- full_join(dfWeight, dfSteps, by = "Date")
> df <- full_join(df, dfCaloriesBurned, by = "Date")
> head(df)
Date Weight Steps CaloriesBurned
1 2016-03-31 74.2 10069 2622
2 2016-03-30 74.9 7688 2538
3 2016-03-29 74.9 4643 2180
4 2016-03-27 74.9 9097 2510
5 2016-03-25 74.3 11160 2777
6 2016-03-24 75.4 8263 2488
I now have tidy data for three Fitbit variables of daily data. Here’s a quick plot of Steps vs Calories burned. 
There’s an obvious relationship since I burn calories with each step. Most of my activities involve making steps – walking, jogging, getting brownies, fleeing swooping birds. I do not clock-up steps when I kayak. I don’t know how Fitbit treats such activities. I’ll check the calorie count before and after next time.
Ultimately I’d like to observe whether any variables can account for outcomes such as weight and sleep. Here’s a Steps vs Weight plot.

There’s no relationship. It remains to be observed whether the inclusion of other Fitbit predictors could account for weight. More on this in future posts.
References and notes
1. I am not affiliated with Fitbit. I’m just a fan. I will however accept gifted Fitbit items if you would like to get in touch with me at willselloutfortech@gmail.com
2. Authenticate with the login function; login(email, password, rememberMe = FALSE). Use your email address and password you use to access your Fitbit dashboard online.
Saturday, 27 June 2015
Probably a better way – Year one
It’s been a year since my first post, and a lot has changed. Looking over my early posts, I’m amused that I started with VBA in Excel. At the time I had completed a VBA training course and was writing a program at work. I saw a future of working exclusively in VBA – writing VB scripts where my customers would use my awfully clunky workbooks to manage and analyse their data. I suspect if I searched “VBA” across job ads, the number of hits has reduced since a year ago.
Soon I commenced the Data Science courses on Coursera. I learnt to program in R via these courses (aided with prior Matlab knowledge) then applied this knowledge at work. I managed my data with R and visualised data with R. I even built a web app (using the Shiny package). R is great, and it’s free. Not like how Facebook is free where one hands over personal data. R is proper free.
I am regularly using R in my new role. I do wonder whether I should learn Python. My understanding is that Python excels over R in web applications and text analytics. I wish I knew Javascript – I would like to create custom interactive displays.
I’d love to know more about statistics and multiple regression. Not entirely sure how I would apply this knowledge. At the heart of it, I’d like to be in a position to receive a large amount of data and simply know what statistical methodology I should be applying towards uncovering insights. I have commenced reading my thick stats book mentioned in this post.
At the moment I am being exposed to different data types and methodologies at work. JSON, XML, MySQL. I’ll take them as they come. I’m fortunate that I am surrounded by developers that have advice when I get stumped. I just have to be more comfortable with asking for help.
Summary: Year one was moving away from VBA to R. Here’s to year two!
Soon I commenced the Data Science courses on Coursera. I learnt to program in R via these courses (aided with prior Matlab knowledge) then applied this knowledge at work. I managed my data with R and visualised data with R. I even built a web app (using the Shiny package). R is great, and it’s free. Not like how Facebook is free where one hands over personal data. R is proper free.
I am regularly using R in my new role. I do wonder whether I should learn Python. My understanding is that Python excels over R in web applications and text analytics. I wish I knew Javascript – I would like to create custom interactive displays.
I’d love to know more about statistics and multiple regression. Not entirely sure how I would apply this knowledge. At the heart of it, I’d like to be in a position to receive a large amount of data and simply know what statistical methodology I should be applying towards uncovering insights. I have commenced reading my thick stats book mentioned in this post.
At the moment I am being exposed to different data types and methodologies at work. JSON, XML, MySQL. I’ll take them as they come. I’m fortunate that I am surrounded by developers that have advice when I get stumped. I just have to be more comfortable with asking for help.
Summary: Year one was moving away from VBA to R. Here’s to year two!
Sunday, 7 June 2015
Get the data – JSON APIs
I have recently been moving away from accessing data via CSVs (comma-separated values) to APIs (application programming interface). An API is “a set of programming instructions and standards for accessing a Web-based software application or Web tool” [1]. Why not just download the CSV then suck it into your analytics tool of choice (mine being R)?
I am developing a new reporting system at work (got me a new job). Automation is a priority. I want users to double click an icon to generate the latest report. Full automation will be achieved by automating the separate process that link one form of data to another. Raw data to tidy data is achieved via my custom R programs. Analysed data to report data could be achieved via a web app accessible by users. These data forms and the process that produce each build the “data pipeline”. I’m not sure if I made up that term myself or I read it online and I’m not attributing the source.
What about just getting the data? The first process in the pipeline it to get the data. Traditionally, I would save the CSV file in a directory, write the script to read the data into R, then run the script. With this approach I have to:
Boom!
However, the setup to download API data took a bit more fiddling-around than I expected.
The survey website API I’m accessing data from permits two flavours – I could download the data as JSON files or XML files. I will not go into detail about how these differ. Look it up yourself. I chose JSON. I actually can’t remember why.
I knew that R had packages to handle JSON files, namely, jsonlite. This is what I needed the package to do:
It took a bit of trial and error to get the download part to work. Eventually the webpage data appeared in R in something that I assumed was JSON format. Here’s a snippet:
I’ll come back to the red box later. The jsonlite package includes a function that converts the JSON format to a data frame. However, after applying it I received an error message indicating a “lexical error”. Something in the JSON format was not in proper JSON format. Unless the data was correctly formatted, I was unable to get my data frame.
Thankfully, I have a colleague who is a JSON guru. His name happens to be Jason. No joke. I’m thinking of changing my name to “R” for consistency. He looked at the format and said it was the “\r\n”. “\r” indicates that there is a new line in the text. The backslash is an “escape character”. The escape character lets R know that the letters “r” and “n” or not actual strings – with the escape characters they become a symbol for a new line.
Jason said I could get past the error by replacing “\r\n” with “\\r\\n”. The double backslash escapes the escape characters. R will read “\\r\\n” as “\r\n\” - a new line. The function gsub finds the old and replaces it with the new
“theJSON” is the string of JSON data. What’s with all the extra escape characters “\\\\r\\\\n”? I require one escape character for each escape character I’m inserting in. Since there are two backslashes before the r and n in “\\r\\n”, I’ll need two more for each - "\\\\r\\\\n".
I re-ran the script and got past the error message. Success! Then I ran into another lexical error. Oh no! Upon each error, I updated the substitutions including:
The updated process:
References and notes
I am developing a new reporting system at work (got me a new job). Automation is a priority. I want users to double click an icon to generate the latest report. Full automation will be achieved by automating the separate process that link one form of data to another. Raw data to tidy data is achieved via my custom R programs. Analysed data to report data could be achieved via a web app accessible by users. These data forms and the process that produce each build the “data pipeline”. I’m not sure if I made up that term myself or I read it online and I’m not attributing the source.
What about just getting the data? The first process in the pipeline it to get the data. Traditionally, I would save the CSV file in a directory, write the script to read the data into R, then run the script. With this approach I have to:
- Visit the location of the data (such as a website)
- Save the data as CSV into a directory
- Run the R script.
- Download the data directly into R.
Boom!
However, the setup to download API data took a bit more fiddling-around than I expected.
The survey website API I’m accessing data from permits two flavours – I could download the data as JSON files or XML files. I will not go into detail about how these differ. Look it up yourself. I chose JSON. I actually can’t remember why.
I knew that R had packages to handle JSON files, namely, jsonlite. This is what I needed the package to do:
- Download the JSON file
- Convert the JSON file format to a data frame (something that resembles a spreadsheet).
It took a bit of trial and error to get the download part to work. Eventually the webpage data appeared in R in something that I assumed was JSON format. Here’s a snippet:
I’ll come back to the red box later. The jsonlite package includes a function that converts the JSON format to a data frame. However, after applying it I received an error message indicating a “lexical error”. Something in the JSON format was not in proper JSON format. Unless the data was correctly formatted, I was unable to get my data frame.
Thankfully, I have a colleague who is a JSON guru. His name happens to be Jason. No joke. I’m thinking of changing my name to “R” for consistency. He looked at the format and said it was the “\r\n”. “\r” indicates that there is a new line in the text. The backslash is an “escape character”. The escape character lets R know that the letters “r” and “n” or not actual strings – with the escape characters they become a symbol for a new line.
Jason said I could get past the error by replacing “\r\n” with “\\r\\n”. The double backslash escapes the escape characters. R will read “\\r\\n” as “\r\n\” - a new line. The function gsub finds the old and replaces it with the new
theJSON <- gsub("\r\n", "\\\\r\\\\n", theJSON)
“theJSON” is the string of JSON data. What’s with all the extra escape characters “\\\\r\\\\n”? I require one escape character for each escape character I’m inserting in. Since there are two backslashes before the r and n in “\\r\\n”, I’ll need two more for each - "\\\\r\\\\n".
I re-ran the script and got past the error message. Success! Then I ran into another lexical error. Oh no! Upon each error, I updated the substitutions including:
- theJSON <- gsub("\t", "", theJSON) # Replace tab symbol
- theJSON <- gsub("[0-9]*\\\\[0-9]*\\\\[0-9]*", 'DATE', theJSON) # Replace date with string "DATE"
The updated process:
- Download the JSON file
- Convert the downloaded data to correct JSON format
- Convert to a data frame.
References and notes
1. http://money.howstuffworks.com/business-communications/how-to-leverage-an-api-for-conferencing1.htm
Wednesday, 12 November 2014
Smart dumb for loops – R
I few weeks ago, I created a monster of a for loop with a bunch of if statements using R. The loop was not doing want I wished it to do, namely, copy some dates into a data frame at indexed positions. And so began my hours of troubleshooting.
I fixed some errors (referencing the wrong variables), rearranged the placement of if statements, fiddled with parentheses and did some general tidying-up. I highlighted the code and ctrl Entered (which runs code in RStudio) and – the errors were gone, yet the dates were not imputed to where they should have been.
I started to question the fundamental way that R for loops operated. Were they so different from the for loops I had used in Matlab and VBA? How deep into the help file would I need to venture to find a solution? Would I need to submit a query on Stack Overflow for the first time?
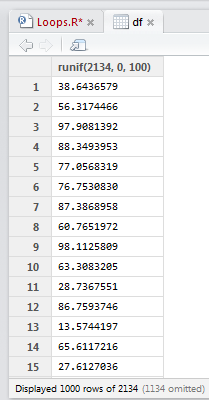
Here's a quick example of what I was dealing with. Take this data frame that has 2134 rows of random numbers between 0 and 100. The start of the data frame is shown.
I fixed some errors (referencing the wrong variables), rearranged the placement of if statements, fiddled with parentheses and did some general tidying-up. I highlighted the code and ctrl Entered (which runs code in RStudio) and – the errors were gone, yet the dates were not imputed to where they should have been.
I started to question the fundamental way that R for loops operated. Were they so different from the for loops I had used in Matlab and VBA? How deep into the help file would I need to venture to find a solution? Would I need to submit a query on Stack Overflow for the first time?
Here's a quick example of what I was dealing with. Take this data frame that has 2134 rows of random numbers between 0 and 100. The start of the data frame is shown.
The code shows a simple for loop that is to iterate through each row and turn each value in an even row to zero. This is achieved using the modulo operation of row number (i) %% 2. If a remainder remains, then the row is odd. No remainder, the row is even.
After running the loop, there is no change to the values in the data frame. WHY?
I don’t know what neuron fired in my head, but it set off a cascade of action potentials that made me want to slap palm against forehead (my palm, my forehead, not someone else's). I had forgotten this.
That's right. A "1" followed by a ":" before nrow(df). Previously I had "i in nrow(df)" assuming that this was sufficient for R to know I wanted it to iterate through each row of the data frame. However, nrow(df) equals the number of rows df, which is 2134. Before, my code read "i in 2134", restricting any changes to the last row alone.
With the correct "i in 1:nrow(df)" the code now reads "i in 1, 2, 3, 4, 5… 2134". The sequential rows of the data frame are specified.
I felt rather stupid and shared this with a friend. His reply:
Ah yes, I know those moments. And I'm never sure if I should feel smart that I figured it out, or dumb that I made the mistake in the first place :)
I settled for a bit of both.
Earlier today a new loop was failing. Turns out I had this.
For the love of pizza – I hadn't even included the "nrow" part around the data frame. That one tipped the scales to the dumb-side.
Sunday, 28 September 2014
Word clouds using the wordcloud package – R
Everyone loves word clouds. But how do you create them? Thankfully, there is an R package for the purpose, called “wordcloud”.
First, I needed to install the wordcloud and the text mining (tm) packages. The RColorBrewer package is required (if you don’t have it already).
I need a bunch of words. I've always liked the introduction speech that V makes in V for Vendetta [1]. I set the words as a character.
I would like to restrict the word cloud to v-words only. There are a number of steps required to present only v words.
Great, but I cannot see the "I" and "you". Such "stop words" are not included by default when using wordcloud. Also notice that "Skyler" and "NASDAQ" are lower case. Further, the apostrophe has been removed from "you're".
First, I needed to install the wordcloud and the text mining (tm) packages. The RColorBrewer package is required (if you don’t have it already).
I need a bunch of words. I've always liked the introduction speech that V makes in V for Vendetta [1]. I set the words as a character.
I would like to restrict the word cloud to v-words only. There are a number of steps required to present only v words.
- The string is split up into separate words as a data frame. Remember, a data frame is "a table, or two-dimensional array-like structure, in which each column contains measurements on one variable, and each row contains one case".
- Get the row numbers of the data frame containing words that start with v. This is achieved by use of metacharacters ^[Vv] which will search for the start of each word with either an upper or lower case v.
- Use the row numbers to subset the words from the data frame. That is, the non v words are excluded.
- Some of these v words end with punctuation marks like full stops and commas. These are removed using the gsub() function and metacharacters [[:punct:]].
- The remaining text of v words free of punctuation are fed into the wordcloud function with parameters defining the size of the text and colours.
Coolness - I have my v words as a word cloud. However this isn't a particularly exciting word cloud since each word only appears once, thus the text are the same size and colours. Let's try the following quote (spoilers!) from Walter White in Breaking Bad [2].
Looking at the quote we can see that the words "I" and "you" occur a number of times. Let's run some code and produce the wordcloud.
Great, but I cannot see the "I" and "you". Such "stop words" are not included by default when using wordcloud. Also notice that "Skyler" and "NASDAQ" are lower case. Further, the apostrophe has been removed from "you're".
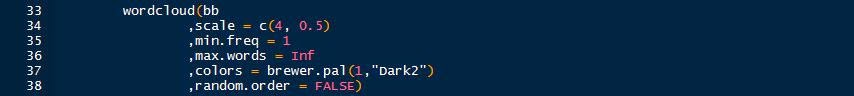
What if I wished to include all words from the quote, adjust the capitals and keep the apostrophes? Starting again with the quote (bb – highlight and run it again), the following code is run [3].
After running the first nine lines, the quote looks like the below.
The code takes the character string, splits it into separate words, counts the frequency of each word, sorts the matrix, creates a data frame that is fed into the workcloud function to produce the word cloud.
Done! All words are present. One could remove the stopwords from the quote by using the following.
Then the code above (in the screengrab before the final Breaking Bad word cloud) can be run to generate a new wordcloud free of the common English words. Happy word clouding!
References and notes
1. Click here for a very excellent V for Vendetta kinetic typography of the speech.
2. If you have yet to watch Breaking Bad, DO NOT watch this spoiler.
3. The keen observers will have noticed bb <- stripWhitespace(bb) as the first line. This command takes the character quote and removed any spaces (such as those created by using tab). This line is required, otherwise the frequency table will tally these white spaces and distort the wordcloud.
Sunday, 13 July 2014
Online R programming course
If I had my time again, I would have taken a gap year before commencing University. Rather than travel the world whilst supporting myself working data entry jobs in windowless offices, I would be particularly nerdy and enrol in a whole bunch of free online courses.
There were very few structured online courses back at the turn of the century. Google was barely a thing.
I didn't know what I wanted to do at University at the age of 17. Yet whilst completing my final year exams, I filled out my list of preferred degrees informed by short blurbs from a phone book-thick course guide. I ended up in a molecular biology/genetics/biochemistry degree and proceeded to hate if for the three year duration. Good choice me [1].
The availability of massive open online courses (MOOCs) allows me to say, "Hey, what's a course in Marine Biology like?", then find one and enrol. In the comfort of my home, I watch videos, attempt quizzes and submit assignments. If I don't understand any of the course content, I can engage with other student on discussion forums [2].
I enrolled and completed two Coursera courses to date: The Data Scientist's Toolbox and R Programming. The courses are really well-structured and the range of topics covered have been broad. R programming included the following:
Course assignments drove the requirement to learn programming in R. One assignment provided hospital mortality data from hundreds of hospitals across the United States. The assignment was to write a function whereby users could enter the name of a US State and request a list that ranked the best (lowest mortality) or worst hospitals (higher mortality). Since I have programming experience, I could list the logical steps required to fulfil the task:
I do know how to code the logical steps via VBA in Excel. However the assignment required coding in R. I had to learn the R syntax required for each step. That is, I had to correctly use the R syntax for a For loop, for an If Else statement, etc. The screen-grab shows a simple For loop with an If Else statement as written in VBA (background) and R (foreground using the R Studio IDE). One step at a time, putting it all together, I had a functioning function.
There were very few structured online courses back at the turn of the century. Google was barely a thing.
I didn't know what I wanted to do at University at the age of 17. Yet whilst completing my final year exams, I filled out my list of preferred degrees informed by short blurbs from a phone book-thick course guide. I ended up in a molecular biology/genetics/biochemistry degree and proceeded to hate if for the three year duration. Good choice me [1].
The availability of massive open online courses (MOOCs) allows me to say, "Hey, what's a course in Marine Biology like?", then find one and enrol. In the comfort of my home, I watch videos, attempt quizzes and submit assignments. If I don't understand any of the course content, I can engage with other student on discussion forums [2].
I enrolled and completed two Coursera courses to date: The Data Scientist's Toolbox and R Programming. The courses are really well-structured and the range of topics covered have been broad. R programming included the following:
- Data types
- Subsetting
- Reading and writing data
- Control structures
- Functions
- Scoping rules
- Vectorized operations
- Debugging.
Course assignments drove the requirement to learn programming in R. One assignment provided hospital mortality data from hundreds of hospitals across the United States. The assignment was to write a function whereby users could enter the name of a US State and request a list that ranked the best (lowest mortality) or worst hospitals (higher mortality). Since I have programming experience, I could list the logical steps required to fulfil the task:
- Read the spreadsheet data
- Point to the State and mortality columns
- Get the user's input parameters (State and rank request)
- Organise the data as specified by the user's parameters (subsetting and sorting)
- Return the result as a list.
I do know how to code the logical steps via VBA in Excel. However the assignment required coding in R. I had to learn the R syntax required for each step. That is, I had to correctly use the R syntax for a For loop, for an If Else statement, etc. The screen-grab shows a simple For loop with an If Else statement as written in VBA (background) and R (foreground using the R Studio IDE). One step at a time, putting it all together, I had a functioning function.
Different programming languages can all perform the same basic tasks (I assume). Efficiency, tools and packages, time, usability, cost, experience, popularity – such factors influence the choice of a programming language. I can see where I would use R for some projects at work over VBA. Then I can enter the debate of whether one should use SAS/SPSS/Python/R for business needs. I think the first two have a greater market share. However the last two are open source and have a dedicated users with various Meetup groups. For now, I'm happy to sample different programming languages via MOOCs. But life still owes me a gap year.
References and notes
1. I believe that it is a tad bit unfair for a 17 or 18 year old to make a choice that significantly dictates the course of one's career for (most of) life. I had had very few major life experiences at 17; I was living at home, was not working, had barely travelled, and only knew how to make plain omelettes and cheese pizzas.
2. A disadvantage to "real life" courses is the lack of direct access to lecturers and tutors. Recognition of qualifications upon completing an online course is under development.
Subscribe to:
Posts (Atom)